Field-Programmable, Recombinase-Based Biomolecular Circuits
This project funded by the Army's Institute for Collaborative Biotechnologies, an Army University Affiliated Research Center (UARC)
|
Project participants:
Additional participants: |
Objectives
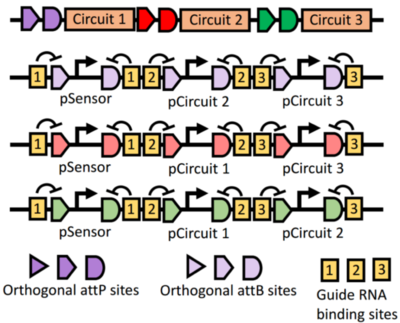
This project explores the use of recombinases -- integrases, excisionases, and other methods of manipulating DNA -- as platform for engineering biomolecular circuits. Our high-level goal is to develop a design-oriented framework for recombinase-based, genetically-encoded circuits that can be used for detection, diagnostics, and logging of environmental signals and events. The specific objectives this project are to:
- Develop circuit components, mathematical models, experimental protocols, and system characterization methods that enable recombinase-based circuits to be designed and utilized in a systematic fashion.
- Implement a set of biological “event detectors” in both cell-free and cell-based assays, and demonstrate their utility in applications that combine digital, analog, and stochastic operations.
References
- Addressable and adaptable intercellular communication via DNA messaging. John P. Marken and Richard M. Murray. Nature Communications, 14:2358, 2023.
- Integrase-mediated differentiation circuits improve evolutionary stability of burdensome and toxic functions in E. coli. Rory L. Williams and Richard M. Murray. Nature Communications, 13:6822, 2022.
- Tunable integrase-mediated differentiation facilitates improved output of burdensome functions in E. coli. Rory L. Williams, Richard M. Murray. bioRxiv 614529, 2019 (revised 2020, 2022).
- Addressable, “Packet-Based” Intercellular Communication through Plasmid Conjugation. John P. Marken, Richard M. Murray. qBio 2019 Conference.
- Proof of concept continuous event logging in living cells. Andrey Shur, Richard M. Murray. 2019 Synthetic Biology: Engineering, Evolution and Design (SEED) Conference.
This research is supported by the Institute for Collaborative Biotechnologies through cooperative agreement W911NF-19-2-0026 from the U.S. Army Research Office. The content of the information on this page does not necessarily reflect the position or the policy of the Government, and no official endorsement should be inferred.
|
|
|