Deciphering the Rules of Nucleus Architecture with Synthetic Cells and Organelles: Difference between revisions
No edit summary |
No edit summary |
||
| (4 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
{{righttoc}} | {{righttoc}} | ||
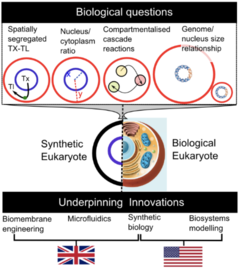
The goal of this | The goal of this project is to decipher the fundamental principles for compartmentalization in cells by building and modeling synthetic cells and organelles from the bottom-up. Working with Imperial College London, we are developing new synthetic biology, microfluidics, biomembrane engineering, and mathematical modelling platform technologies. These are being used to make and model synthetic nuclei that will be used to address fundamental questions relating to its central role in all eukaryotes, by taking an "understanding by building" approach. This research aims to establish a new framework for understanding cell biology fundamentals, that can be used in future studies by us and others. By the end of the project, we will have developed a technological toolkit to probe the principles underpinning compartmentalization in cells, and use these to answer key unanswered biological questions that are uniquely placed to be addressed using our approach. | ||
{| cellpadding=0 cellspacing=0 width=80% | {| cellpadding=0 cellspacing=0 width=80% | ||
| Line 11: | Line 11: | ||
| | | | ||
Collaborators: | Collaborators: | ||
{{project collaborators}} | |||
Past participants: | Past participants: | ||
| Line 19: | Line 18: | ||
=== Objectives === | === Objectives === | ||
[[Image:bbsrc-synnucleus.png|right| | [[Image:bbsrc-synnucleus.png|right|240px]] | ||
Caltech's activities in this collabrative proposal with Imperial College London are focused on the following objectives: | Caltech's activities in this collabrative proposal with Imperial College London are focused on the following objectives: | ||
* Development of model-based analysis, design and systems optimization for synthetic organelles. This work will leverage prior work in open-source software tools for modeling of biomolecular systems and will result in extended modeling and analysis tools that will be made available for distribution both to the project and the public. | * Development of model-based analysis, design and systems optimization for synthetic organelles. This work will leverage prior work in open-source software tools for modeling of biomolecular systems and will result in extended modeling and analysis tools that will be made available for distribution both to the project and the public. | ||
* Systematic exploration of the effects of compartmentalization on coupled biochemical processes, using the experimental platforms developed at Imperial to validate and build on modeling predictions. | * Systematic exploration of the effects of compartmentalization on coupled biochemical processes, using the experimental platforms developed at Imperial to validate and build on modeling predictions. | ||
=== References === | === References === | ||
{{project paper list}} | {{project paper list}} | ||
[[Category: | [[Category:Completed projects]] | ||
[[Category:Biocircuits projects]] | [[Category:Biocircuits projects]] | ||
{{Project | {{Project | ||
Latest revision as of 17:31, 5 October 2024
The goal of this project is to decipher the fundamental principles for compartmentalization in cells by building and modeling synthetic cells and organelles from the bottom-up. Working with Imperial College London, we are developing new synthetic biology, microfluidics, biomembrane engineering, and mathematical modelling platform technologies. These are being used to make and model synthetic nuclei that will be used to address fundamental questions relating to its central role in all eukaryotes, by taking an "understanding by building" approach. This research aims to establish a new framework for understanding cell biology fundamentals, that can be used in future studies by us and others. By the end of the project, we will have developed a technological toolkit to probe the principles underpinning compartmentalization in cells, and use these to answer key unanswered biological questions that are uniquely placed to be addressed using our approach.
|
Current participants: Additional participants: |
Collaborators:
Past participants:
|
Objectives
Caltech's activities in this collabrative proposal with Imperial College London are focused on the following objectives:
- Development of model-based analysis, design and systems optimization for synthetic organelles. This work will leverage prior work in open-source software tools for modeling of biomolecular systems and will result in extended modeling and analysis tools that will be made available for distribution both to the project and the public.
- Systematic exploration of the effects of compartmentalization on coupled biochemical processes, using the experimental platforms developed at Imperial to validate and build on modeling predictions.
References
- Impact of Chemical Dynamics of Commercial PURE Systems on Malachite Green Aptamer Fluorescence. Zoila Jurado, Richard M. Murray. Submitted, ACS Synthetic Biology, 2024.
- Optimizing protein expression in the One-Pot PURE system: Insights into reaction composition and translation efficiency. Y. Zhang, Y. Qiu, M. Deveikis, Z. A. Martinez, T.-F. Chou, P. Freemont, R. M. Murray. SEED, 2024.
Research supported by the National Science Foundation award number 2152267.
|
|
|