Multi-Layer, Composable and Programmable Biomolecular Circuits for Microbial Consortia: Difference between revisions
No edit summary |
|||
| Line 17: | Line 17: | ||
=== Objectives === | === Objectives === | ||
[[Image:icb-multilayer.png|right| | [[Image:icb-multilayer.png|right|240px]] | ||
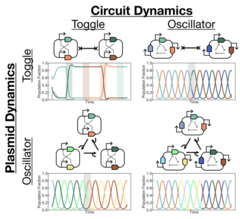
Our objective for this project is to demonstrate that by decoupling transfer blocking from plasmid cleavage, we can engineer programmable behavior that operates at the population-level layer. Then, by encoding variants of a given transcriptional circuit on different plasmid types, we will demonstrate the composability of circuit dynamics across multiple layers. Finally, we will construct strain-level circuit elements and compose them with circuit elements at the transcriptional and plasmid layers to demonstrate how multilayer systems lead to multiplicative scaling in the number of system configurations. | Our objective for this project is to demonstrate that by decoupling transfer blocking from plasmid cleavage, we can engineer programmable behavior that operates at the population-level layer. Then, by encoding variants of a given transcriptional circuit on different plasmid types, we will demonstrate the composability of circuit dynamics across multiple layers. Finally, we will construct strain-level circuit elements and compose them with circuit elements at the transcriptional and plasmid layers to demonstrate how multilayer systems lead to multiplicative scaling in the number of system configurations. | ||
=== References === | === References === | ||
{{project paper list}} | {{project paper list}} | ||
Revision as of 17:33, 11 December 2021
The long-term goal of this project is to create the mathematical, experimental, and application framework for recombinase-based biomolecular circuits, with a new focus on distributed operations in microbial consortia. Initial work has demonstrated the construction of a synthetic plasmid conjugation system that can pass plasmids to defined recipient strains within a population. In this project, we will build on those results and use plasmid conjugation as a means to implement multi-strain engineered microbial consortia with dynamic properties that are relevant for a range of modular, composable and programmable biological behaviors, including event logging, materials synthesis, programmable biofilms, and engineered living materials.
|
Current participants: Additional participants: |
Collaborators: Past participants:
|
Objectives
Our objective for this project is to demonstrate that by decoupling transfer blocking from plasmid cleavage, we can engineer programmable behavior that operates at the population-level layer. Then, by encoding variants of a given transcriptional circuit on different plasmid types, we will demonstrate the composability of circuit dynamics across multiple layers. Finally, we will construct strain-level circuit elements and compose them with circuit elements at the transcriptional and plasmid layers to demonstrate how multilayer systems lead to multiplicative scaling in the number of system configurations.
References
None to date
This research is supported by the Institute for Collaborative Biotechnologies through cooperative agreement W911NF-19-2-0026 from the U.S. Army Research Office. The content of the information on this page does not necessarily reflect the position or the policy of the Government, and no official endorsement should be inferred.
|
|
|