SURF 2017: Integrase based circuits for sequential and temporal control of gene expression
2017 SURF: project description
- Mentor: Richard M. Murray
- Co-mentors: Rory Williams
Integrase based logic gates rely on the ability of integrases to invert, excise, or integrate functionally important sequences of DNA flanked by pairs of integrase recognition sites. Using both orthogonal integrases as well as multiple orthogonal integrase attachment sites per integrase, two-input one-output logic gates, two and three input state-machines, as well as temporal logic gates have been successfully demonstrated (Bonnet et al., 2013; Roquet et al., 2016; Hsiao et al., 2016). However, little has been done to harness integrase based logic for the control of gene expression in a sequential manner. This project would explore the utility of integrases for controlling sequential gene expression, and determine how to control the timing of steps in the cascade through tuning of promoter strength, mRNA stability, degradation tags, plasmid copy number, and/or other parameters. An example circuit for sequential gene expression is shown below to demonstrate the general principle, though the student may design a different circuit for this project. This project would involve modeling of the circuit to determine desirable characteristics and narrow the parameter space, and assembly and testing of the circuit(s) in vitro in TX-TL and in vivo in cells.
Recommended skills: Students interested in this project should have some basic laboratory skills (equivalent to Bi 1X, Bi 10 or ChE/BE 130) and have experience with either MATLAB, Python or a similar scientific programming package.
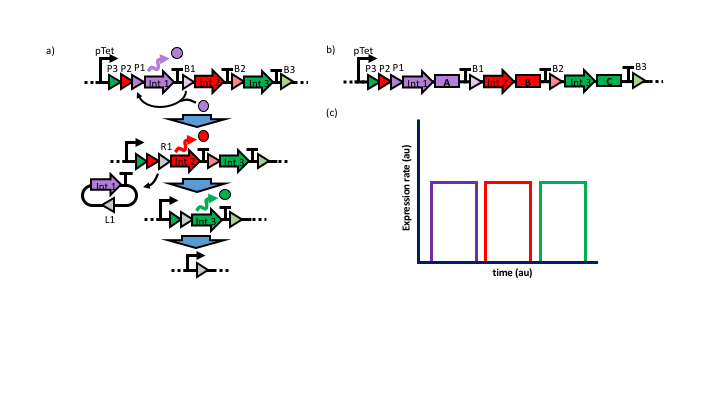
Figure 1: (a) The circuit is constructed to have a single inducible promoter in front of an array of integrase genes. Each integrase coding sequence is followed by a transcriptional terminator and flanked by parallel attB/attP sites. Expression of the furthest upstream integrase results in the excision of its corresponding coding sequence, and allows expression of the subsequent integrase. (b) Incorporation of polycistronic transcripts in place of the single integrase allows sequential, non-overlapping expression of desired genes or operons (c).
References
J. Bonnet, P. Yin, M. E. Ortiz, P. Subsoontorn, and D. Endy. Amplifying genetic logic gates. Science, 340(6132):599–603, 2013.
V. Hsiao, Y. Hori, P. W. K. Rothemund, and R. M. Murray. A population-based temporal logic gate for timing and recording chemical events. Molecular Systems Biology, 12(5):869, 2016.
N. Roquet, A. P. Soleimany, A. C. Ferris, S. Aaronson, and T. K. Lu. Synthetic recombinasebased state machines in living cells. Science, 353(6297):aad8559, 2016.