SURF 2015: Machine-readable protocols and rapid-prototyping for synthetic biology research
2015 SURF project description
- Mentor: Richard M. Murray
- Co-mentor: Scott C. Livingston and Sean Sanchez

One of the pillars of science is repeatability. For synthetic biology, repeatability is also important because it facilitates the composition of multiple methods to yield new (combined) circuits in vitro and in vivo. Stated broadly, a central concern of synthetic biology is predictable engineering of chemical processes within cells, and a crucial enabler to this end is repeatable experiments. Despite methodological progress within particular labs or by particular people, sharing protocols that work "out-of-the-box" remains challenging. There has been some effort to share and organize protocols [1].
We are thus motivated to develop a machine-readable format for protocols. By "machine-readable" we mean one that may be reliably parsed by software. Such a format would provide the basis for several improvements to current practices:
- scheduling to manage completion of steps of a protocol and negotiate collaborations and triage during interruptions;
- assessing capabilities of lab equipment to decide whether it can be used for some steps;
- automating steps when the approriate tool is available, supporting manual tasks when not;
- facilitating composition of protocols;
- versioning (tracking the history of modifications).
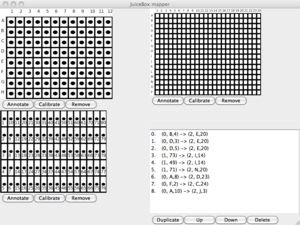
Toward achieving these, a summer research project would focus on the construction of a tool that automates some parts of protocols using TX-TL [2] and provides a platform on which to explore formats for experiment protocols that could realize the above ambitions. Throughout the summer, there would be an emphasis on agile design practices, in which the student will make use of rapid-prototyping technologies, like so-called 3D-printing. While the project could focus on a different tool, some initial progress has been made toward a liquid-handling robot (Figures 1 and 2) based on the MTM Snap milling machine.
While there will be involvement with traditional activities like pipetting in a cell biology lab, much of this project will focus on engineering. Though not necessary, it would be useful for the student to have some knowledge about one or more of the following:
- electronic circuit layout; e.g., using a CAD tool like EAGLE
- motors of the scale and type used on small RC vehicles
- mechanical design using CAD tools like SolidWorks
- comfortable at the UNIX terminal
- Python or C programming language
- GUI development; e.g., as in Web design or desktop applications
The nature of the project is pliable according to skills with which the student arrives. So, no need to worry if you only know about one or two of the above!
References
[1] OpenWetWare from the BioBricks Foundation
[2] Z.Z. Sun, C.A. Hayes, J. Shin, F. Caschera, R.M. Murray, V. Noireaux (2013). Protocols for Implementing an Escherichia coli Based TX-TL Cell-Free Expression System for Synthetic Biology. J. Vis. Exp. (79), e50762, DOI: 10.3791/50762