SURF 2015: Design space exploration of the violacein pathway in TX-TL: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 1: | Line 1: | ||
'''[[SURF 2015|2015 SURF]] project description''' | |||
*Mentor: Richard Murray | *Mentor: Richard Murray | ||
*Co-mentor: Shaobin Guo and Yong Wu | *Co-mentor: Shaobin Guo and Yong Wu | ||
Revision as of 20:23, 19 December 2014
2015 SURF project description
- Mentor: Richard Murray
- Co-mentor: Shaobin Guo and Yong Wu
Project Description
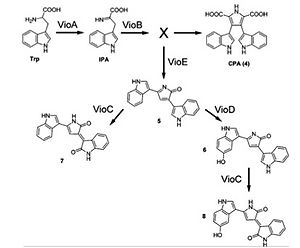
Violacein is a water-insoluble violet pigment that naturally exists in a few bacteria (Jiang, 2010 & Pantanella, 2007). Violacein has the potential applications in antibacterial, anti-trypanocidal, anti-ulcerogenic, and anticancer drugs (Balibar, 2006). The pathway to produce violacein from tryptophan consists of five enzymes vioA-E (Hoshino, 2011). The pathway is NADPH dependent and consumes oxygen. The various enzymes involved in the pathway could also lead to the production of a few colorful byproducts, which might interfere with fluorescence signals when plate reader is used. The goal of the project is to characterize the violacein pathway using cell-free transcription and translational (TX-TL) system (Sun, 2013). Student will take advantage of the TX-TL system to explore ways to improve violacein production.
References:
Balibar, C. J. and C. T. Walsh (2006). "In Vitro Biosynthesis of Violacein from l-Tryptophan by the Enzymes VioA−E from Chromobacterium violaceum." Biochemistry 45(51): 15444-15457. [1]
Hoshino, T. (2011). "Violacein and related tryptophan metabolites produced by Chromobacterium violaceum: biosynthetic mechanism and pathway for construction of violacein core." Applied Microbiology and Biotechnology 91(6):1463-1475.
Jiang, P.-x., et al. (2010). "Reconstruction of the violacein biosynthetic pathway from Duganella sp. B2 in different heterologous hosts." Applied Microbiology and Biotechnology 86(4): 1077-1088.
Pantanella, F., et al. (2007). "Violacein and biofilm production in Janthinobacterium lividum." Journal of Applied Microbiology 102(4): 992-999.
Sun, Z. Z., et al. (2013). "Protocols for Implementing an Escherichia coli Based TX-TL Cell-Free Expression System for Synthetic Biology." (79): e50762.
Sun, Z. Z., et al. (2013). "Linear DNA for Rapid Prototyping of Synthetic Biological Circuits in an Escherichia coli Based TX-TL Cell-Free System." ACS Synthetic Biology.